INTRODUCTION
Rice is one of the most important cereal
crops and provides the staple food for about half of the world’s population (Moosavi et al., 2015). Genetic enhancement is one of the
important tools to improve the productivity. (Babu et al., 2012). Knowledge on the nature and
magnitude of the genetic variation governing the inheritance of quantitative
character like yield and its components is essential for effecting genetic
improvement. (Rakesh et al., 2015).
A critical analysis of the genetic variability parameters, namely, genotypic
coefficient of variability (GCV), phenotypic coefficient of variability (PCV),
heritability and genetic advance for different traits of economic importance is
a major pre-requisite for any plant breeder to work with crop improvement programmes. (Kishore et al., 2015).
Genetic variability for agronomic traits is the key component of breeding
programs for broadening the gene pool of rice and other crops. The genetic
coefficient of variation together with heritability estimate would give the
best picture of the amount of advance to be expected from selection. The amount
of genetic advance under selection depends mainly on the amount of genetic
variability. (Immanuel et al., 2011).
Development of high
yielding varieties requires the knowledge of existing genetic variability.
Hence, rice breeders are interested in developing varieties with improved yield
and other desirable agronomic characters. (Idris and Mohamed, 2013). Most agronomically
significant characters are inherited quantitatively and are known to be
affected by environmental factors. Selection based on the phenotype would be
difficult for such difficult traits. In breeding programs, it is often
difficult to manipulate such traits, since several inter-componential characters indirectly control them. (Immanuel
et al., 2011).
Genetic variability among traits is important
for breeding and in selecting desirable types. The low heritability of grain
characters made selection for high yielding varieties possible usually using
various components traits associated with yield. (Mulugeta et al., 2012). High magnitude of
variability in a population provides the opportunity for selection to evolve a
variety having desirable characters. (Bornare
et al., 2014). Therefore, the objectives of this study is to determine
the genetic variation and diversity analysis of two hundred and twenty nine
rice genotypes and an improved Nigerian variety (FARO44) were evaluated based
on agronomic traits.
MATERIALS
AND METHODS
The experiment was conducted at experimental
field of African Rice Centre at International Institute of Tropical Agriculture
(IITA), Old Oyo road, Ibadan, Oyo state, Nigeria. During 2020
cropping season. IITA is located at longitude 7º30’8’’N, latitude
3º54’37’’E and at elevation 243m above sea level (Ariyo et al., 2018). The experimental materials of
this study consisted of two
hundred and thirty-nine (239) accessions of rice anther culture derived from
South Korea, of O. sativa L. and one improved and adapted Nigeria rice
variety (FARO 44) collected from African Rice Center, IITA used as check. The
experiment was established in alpha lattice design (rows × column) with four
columns and planted sixty entries in each column (4×60) with two replications.
The seedlings were
raised in the wet prepared nursery bed and then transplanted to the field after
25 days under irrigated system. A single row with size of 0.2×3 m was used as
plot. A single seedling was transplanted per hill at the spacing of 20×20 cm
between rows and between hills. The chemical fertilizer NPK (15:15:15) was
applied as a basal application of 200 kg/ ha (N2, P2O5 and K2O). Urea was applied at the rate of 65 kg/ha at tillering stage and the second rate of 35 kg/ha was applied
at the beginning of panicle initiative (booting) stage. The weeds were controlled twice by using selective herbicide Vespanil Plus (250 ml/ 20 liters of water) at early stage
of crop development and before flowering.
Data was collected at
appropriate stage of the crop development. The agronomic characters were
measured by randomly selected plants from each experimental unit (row). The
Standard Evaluation System (SES) by International Rice Research Institute
(IRRI) for Rice reference manual IRRI, 2002 was used for all trait measurements
except where stated otherwise. The data collected were plant height (cm),
number of tillers per plant, days to 50% flowering, flag leaf length (cm),
panicle fertility (%), productive tillers per plant, panicle length (cm), panicle
weight (g), biomass (g), number of grains per panicle, 1000 grain weight (g),
grain yield per plant (g) and grain yield per plot (kg/ha). Leaf area (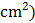 was measured manually following the formula:
was measured manually following the formula:
LA=W×L×K
Where:
LA≡ Leaf Area, W ≡ Leaf Width, L ≡ Leaf length and K ≡
Constant = 0.75
Leaf area index ( ) was
calculated as described (Efisue and
Dike,
2020) as follow:
) was
calculated as described (Efisue and
Dike,
2020) as follow:
LAI = LA / area covered by the plant
Where: LAI ≡ Leaf Area Index, LA ≡
Leaf Area
Genetic parameters were estimated to identify
genetic variation among accessions and to determine genetic and environmental effects
on various characters. These genetic parameters were calculated as described
(Burton and Devane, 1953 and Johnson et al., 1955).
Genotypic variance: 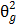 = MSG-MSE/r
= MSG-MSE/r
Where MSG is the mean square of genotypes,
MSE is mean square of error, and r is number of replications
Phenotypic variance:
 =
= 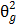 ×
× 
Where  is
the genotypic variance and
is
the genotypic variance and  is the mean squares of error.
is the mean squares of error.
Genotypic coefficient of variance (GCV) and
phenotypic coefficient of variance (PCV):
GCV (%) = √ /
/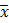 × 100
× 100
PCV (%) = √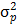 /
/ 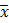 × 100
× 100
Where  = Genotypic variance,
= Genotypic variance,  = Phenotypic variance,
= Phenotypic variance, 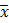 = grand mean of the trait.
= grand mean of the trait.
GCV and PCV values were categorized as low
(0-10%), moderate (10-20%) and high (20% and above).
Heritability (Broad sense) was computed as
described (Amegan et al.,
2020).
 =
=  × 100
× 100
Where  is the genotypic variance and
is the genotypic variance and 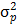 is the phenotypic variance
is the phenotypic variance
The Heritability was categorized as low
(0-30%), moderate (30-60%) and high (60% and above) as described by Robinson et
al, (1949).
Expected
genetic advance (GA):
Expected genetic advance (GA) was computed
according to (Allard, 1960) cited by (Saeed et
al., 2018) as:
GA =  ×
× 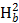 × k
× k
Where, GA = Expected genetic advance, = Phenotypic standard deviation,
= Phenotypic standard deviation, 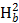 = Heritability in broad sense
= Heritability in broad sense and k =
the standardize selection differential at 5% selection intensity (K = 2.063).
and k =
the standardize selection differential at 5% selection intensity (K = 2.063).
Principal component analysis was computed by
using the principal component procedure of multivariate technique in
Statistical Analysis System SAS, version 9.4 (2018) for analyzing data for all
the characters. Cluster analysis was carried out to clustering the genotypes
into different groups using R software version 3.9.6.
RESULTS AND DISCUSSION
Genetic Variability parameters
High
variability in the initial breeding material ensures better chances of
producing desired crop plant. Genotypic coefficient of variation (GCV) measures
the variability of any trait. The extent of the environmental influence on any
trait is indicated by the magnitude of the differences between the genotypic
and phenotypic coefficients of variation. Large differences reflect high
environmental influence as reveal by high phenotypic influence, while small
differences reveal high genetic influence (Idris
and Mohamed, 2013). In this study phenotypic
coefficients of variance were higher than the corresponding genotypic
coefficients of variance for all the studied traits (Table 1). This indicates
the presence of environmental influence to some degree in the phenotypic
expression of the characters (Iftekharuddeula et
al., 2001 and Idris et al., 2012) observed
similar results. Likewise (Singh et al., 2019, Sameera et al., 2016, Rakesh et al., 2015 and Mulugeta et al., 2012).
The differences were observed between genotypic and phenotypic
coefficients of variance for days to 50% flowering, plant height, panicle
length, 1000 grain weight and fertility (Table 1). This indicates that
genotypic influence had played an important role rather than phenotypic
influence indicating less influence of environment on these traits. Therefore,
selection on the basis of phenotype alone can be effective for the improvement
of these traits (Idris and Mohamed, 2013). High
differences between genotypic and phenotypic coefficients of variance were
observed for biomass, grain yield, grain yield per
plant, number of grains per panicle, panicle weight, and effective tillers.
Therefore, indicating that phenotypic influence assert more on the existence of
environment influence on these traits.
The
characters studied in the present investigation exhibited high, moderate to low
PCV and GCV values. The high PCV (>20%) and GCV values were recorded for
grain yield, grain yield per plant, biomass, number of tillers, panicle weight,
effective tillers, leaf area, leaf area index and number of grains per panicle
(Table 1). The result in conformity with the reports of (Kishore et al., 2015). for
grain yield; (Prasad et al., 2013) for number of tillers per plant; (Sameera et al., 2016) for
number of effective tillers and number of grains per panicle; (Hasib et al., 2004) for grain yield per plant; (Bekele et al., 2013) for effective tillers per
plant; (Bornare et al., 2014) for effective tillers and grain yield per
plant; (Singh et al., 2019)
also recorded similar observation for grain yield per plant and number of
tillers. Hence, selection on the basis of these phenotypic characters in these
genotypes can also be effective for improvement of grain yield. However,
moderate (10-20%) genotypic and phenotypic coefficients of variance were
recorded in the present study for plant height (PH), days to 50% flowering
(D50%F), panicle length (PL) and 1000 grain weight (TGW) (Table 1). These
results corroborate the earlier reports (Singh et al., 2019) for days to 50% flowering, panicle weight
and 1000 grain weight and for days to 50% flowering (Bornane
et al., 2014 and Kishore et al., 2015). In contrast, low (<10%) estimates of
genotypic and phenotypic coefficients of variation were observed in fertility
(FER).
Heritability
The heritability estimates act as predictive instrument in expressing
the reliability of phenotypic value. Therefore, good knowledge of transmission
of a particular trait assists the plant breeders in predicting the behavior of
succeeding generations because high heritability of a trait simplifies the
selection procedure (Khaliq et al., 2009). In this
study, heritability in broad sense was calculated for all characters under
study and is presented in (table 1). Heritability was classified as low
(<30%), medium (30-60%) and high (>60%). High heritability was recorded
for days to 50% flowering (80.07), plant height (79.65), 1000 grain weight
(75.44), panicle length (74.93) and number of tillers (62.38). These results
are in conformity with the reports of (Idris et al.,
2013) for 1000 grain weight and plant height; (Bisne
et al., 2009) for number of tillers; (Dhanwani et
al., 2013) for days to 50% flowering. Similar results were also quoted by (Rakesh et al., 2015), (Subbaiah et al.,
2011) and Ananadarao et al., 2011). Low
heritability value was recorded for fertility (20.24). High heritability values
indicate that the characters under study are less influenced by environment in
their expression, therefore, the selection may base on the phenotypic
expression of these characters in the individual plant by adopting simple
selection methods. High heritability indicates the scope of genetic improvement
of these characters through selection. The low heritability recorded for
fertility indicate that direct selection for this
trait will be ineffective.
Genetic
advance
The
genetic advance is a useful indicator of the progress that can be expected as
result of exercising selection on the pertinent population. Genetic advance I (Table 1) was highest for grain yield . This result is in conformity with the findings of (Kishore et al., 2015, Dhanwani et al., 2013 and Bornare et al., 2014). This suggest that this
character is predominantly controlled by additive gene action. Hence genetic
improvement through selection for this trait may be effective
(Singh et al., 2019). The
lowest genetic advance was noticed for leaf area index (3.188).
The
information on genetic variation, heritability and genetic advance helps to
predict the genetic gain that could be obtained in later generations, if
selection is made for improving the particular trait under study. Selection for the traits having high heritability
coupled with high genetic advance is likely to accumulate more additive genes
leading to further improvement of their performance (Panse
and Suhatme, 1957) cited by (Rakesh et al., 2015).
High heritability along with high genetic advance was noticed for traits days
to 50% flowering and plant height (Table 1) this was also reported by (Seyoum et al., 2012 and Rakesh et al., 2015). Characters which showed high heritability coupled with
moderate or low genetic advance such as 1000 grain weight, panicle length and
number of tillers can be improved by intermating
superior genotypes of segregating population developed from combination
breeding (Samadhia, 2005), (Babu et al., 2012).
Cluster
Analysis of Rice Genotypes Evaluated
Cluster
analysis of some agronomic traits and yield was conducted and dendrogram was constructed using rice genotypes values
presented in the (Figure 1). Four major groups (A, B, C, and D) were observed
among 240 rice genotypes including an improved variety (FARO44). Cluster analysis provide a useful means for estimating morphological
diversity within and between genotypes evaluated. Its useful
tool to detect the potential of breeding value. Cluster A consisted of
73 genotypes, Cluster B had least genotypes (20) of the group with two sub
groups; B1 contain three genotypes (UPN_643, UPN_611 and UPN_639) and B2
comprise 17 genotypes. Cluster C consisted 35 genotypes.
Cluster D recorded 112 genotypes which is the biggest and the check (FARO44)
was clustered in this group (Figure 1) and FARO 44 contains fortified iron (Ikuli et al., 2017) that showed more genetic relatedness
among the genotypes as compared to any other cluster. Cluster
analysis showed the genetic
variation among 240 genotypes
indicating wide diversity of genetic material. Hence, crossing between two
groups could give
good result and cluster also assist breeders in making good choice in selection
of parents in crop improvement. However, breeders need to evaluate genotypes
accurately in each grouping before use in rice breeding programme
(Maji et al., 2012). For the
selection of parents, genetic diversity is one of the important decisive
factors that will enhance crop improvement (Tuhina-khatun et al., 2015).
Principal
Components Analysis
The result
of principal component analysis explained the genetic diversity of the rice
accessions for all traits under study. The PCA with eigenvalues >1 and which
explained 20% of the variation were considered. Out of fifteen only three
principal components exhibited more than one eigenvalue, and showed 64.78% of
total cumulative variability among the traits studied (Table 2). It also revealed
that the first principal component accounted for 37.17% of total variance and
traits such as plant height (0.32), panicle weight (0.32), panicle length
(0.31), biomass (0.31) and grain yield per plant (0.30) are the most positively
contributed. The second
component accounted for 20.39% of the total variance with most positively
contribution variables such as effective tillers (0.37), number of tillers per
plant (0.31) and grain yield (0.30). The third principal component accounted
for 7.22% and the variables contributing most positively were number of tillers
(0.55), biomass (0.39) and effective tillers (0.32). These results conformed
presence of strong differences among genotypes. Through the PCA, the number of
characters, which are responsible for the observed variation within a group could be identified. The principal component with
higher eigenvalues and variables which had high loading were considered as best
representative of system attributes (Pachauri et
al., 2017), which
may be considered in the utilization of these traits for crop improvement.
These result in agreement with the findings of (Ayenew et al., 2020)
and (Worede et al., 2014) explained 73.5% of the
total variability and (Singh et al., 2013)
explained 62.72% of total variation in rice using the first three principal
components.
Agronomic
parameters projection in the PCA plots showed the phenotypic variation among
the populations this indicated how dispersed along each principal component
(Figure 2). The genotypes UPN540 and UPN632 on the top right part of the plot
showed the highest performance in either axis of principal component 1 and 2. While the genotypes UPN643 is the lowest
performance in either axis of the principal component 1 (Figure 2). The first
three principal components accounted for 64% of the total variation, which
indicated a very strong correlation among the characters being studied.
Accordingly, in the first PCA, the plant height, panicle length, biomass,
panicle weight, and grain yield per plant were important in separating the
genotypes due to their high loadings. Similarly, (Tuhina-khatun et al., 2015 and Worede et al.,
2014) explained 61.2% of the total variability using the first and second
principal components.
CONCLUSION
Results of
variability indicated that there is adequate genetic variability among the genotypes
studied. Characters such as grain yield per plant, biomass and number of
tillers showed high genotypic and phenotypic coefficient variance whereas days
to 50% flowering, plant height, panicle length and 1000 grain weight showed
high heritability likewise number of grains per panicle, biomass, leaf area,
days to 50% flowering and plant height showed high genetic advance suggesting a
scope for improvement of grain yield through selection. Dendrogram
cluster analysis based on quantitative traits classified 240 rice genotypes
including an improved variety (FARO44) into four groups (A, B, C and D) which
detected the divergences among genotypes. The principal component analysis
revealed that the first three components (PC1, PC2 and PC3) accounted for 64.78%
of the total cumulative variability among the traits studied confirmed the
presence of ample genetic diversity for use in improvement.
Acknowledgement
Authors wish to express their gratitude to KAFAC of RDA Korea for
providing the genetic materials (anther culture
derived) used for this study under the project KAR20190112. Gratitude
to Pan African University, Life and Earth Science Institute (PAULESI) for
granting Salim Hassan Kafi scholarship
for this study.
REFERENCES
Allard, R. W. 1960. Principles of Plant
Breeding. John Wiley and Son Inc., New York, USA.
485
Amegan, E., Efisue, A., Akoroda, M., Shittu, A. and Tonegnikes, F. 2020. Genetic Diversity of Korean Rice (Oryza
Sativa L.) Germplasm for Yield and Yield
Related Traits for Adoption in Rice Farming System in Nigeria. International
Journal of Genetics and Genomics 8.1:19–28.
https://doi.org/10.11648/j.ijgg.20200801.13
Ananadrao, S. D., Singh, C. M.,
Suresh, B. G. and Lavanya, G. R. 2011. Evaluation of rice hybrids for yield and yield component characters
under North East Plain Zone. The Allahabad farmer.
67.1:63–68.
Ariyo, O. C., Okojie, L. O., and Ariyo, M. O. 2018. Villagers Willingness to Pay for Forest Conservation in Ibadan, Oyo
State, Nigeria. Asian Journal of Agricultural Extension, Economics
& Sociology 23.4:1–14. https://doi.org/10.9734/AJAEES/2018/40142
Ayenew, A., Dejene, T., and Worede, F. 2020.
Genetic divergence analyses of lowland rice genotypes in North Western
Ethiopia. African Journal of Plant Science 14.4:165–171.
https://doi.org/10.5897/AJPS2019.1782
Babu,
V. R., Shreya, K., Dangi,
K. S., Usharani, G. and Nagesh,
P. 2012. Genetic Variability Studies for Qualitative and Quantitative traits in
Popular Rice (Oryza sativa L.) Hybrids
of India. International Journal of Scientific and Research
Publications 2.6:1–5.
Bekele, B. D., Rakhi, S., Naveen, G. K.,
Kundur, P. J. and Shashidhar,
H. E. 2013. Estimation of genetic variability and correlation studies for grain
zinc concentrations and yield related traits in selected rice (Oryza sativa L.) genotypes. Asian J.
Expt. Bio. Sci. 4.3:391–397.
Bisne, R., Sarawgi, A. K. and Verulkar, S. B. 2009. Study of heritability, genetic advance and variability for yield
contributing characters in rice. Bangladesh Journal of Agricultural
Research 34.2:175–179.
Bornare, S. S., Mittra, S. K. and Mehta A. K. 2014. Genetic
Variability, Correlation and Path Analysis of Floral, Yield and its Component
Traits in CMS and Restorer Lines of Rice (Oryza
sativa L.). Bangladesh J 43.1:45–52.
Burton, G. W. and Devane, E. H.
1953. Estimating heritability in tall fescue (Festuca
arundinacea) from replicated clonal material. Agronomy
Journal 45:478–481.
Dhanwani, R. K., Sarawgi, A. K., Akash. S. and A. Jitendra,
K. T. 2013. Genetic Variability Analysis for Various Yield Attributing
and Quality Traits in Rice (O. sativa L.). The Bioscan
- International Quarterly Lournal of Life Sciences
8.4:1403–1407.
Efisue, A. A., and Dike, C. C. 2020. Screening Rice (Oryza sativa L) for
Salinity Tolerance for Yield and Yield Components in Saline Stressed
Environment 8.1:15–21.
https://doi.org/10.11648/j.ajaf.20200801.13
Hasib, K. M., Ganguli, P. K. and Kole, P. C. 2004.
Evaluation of the performance of advanced generation lines of mutant x Basmati
crosses of scented rice. J. Inter Academicia 8.1:7–10.
Idris, A. E. and Mohamed, K. A. 2013. Estimation of Genetic Variability and Correlation for Grain Yield
Components in Rice. Global Journal of Plant Ecophysiology
3.1:1–6.
Idris, A. E., Justin, F. J., Dagash, Y.
M. I. and Abuali, A. I. 2012. Genetic Variability and
Inter Relationship between Yield and Yield Components in Some Rice Genotypes. American Journal of Experimental Agriculture. 2.2:233–239.
Iftekharuddeula, M., S. Hassan, M. J., Islam, M.A., Badshah,
M.R. and Islam, A. K. 2001. Genetic evaluation and selection criteria of Hybrid rice in
irrigated ecosystem of Bangladesh. Pak. J. Bio. Sci. 4.7:790–791.
Immanuel, S. C., Nagarajan, P., Thiyagarajan, K., Bharathi, M.,
and Rabindran, R. 2011.
Genetic parameters of variability, correlation and path- coefficient studies
for grain yield and other yield Attributes among rice blast disease resistant
genotypes of rice (Oryza sativa L.). African
Journal of Biotechnology 10.17:3322–3334. https://doi.org/10.5897/AJB10.2575
IRRI. 2002. “Standard Evaluation System for Rice.” Find out how
the qualities of rice are evaluated and scored in authoritative sourcebook.
Johnson, H.W., Robinson, H. F. and Comstock, R. E.
1955. Estimates of genetic and environmental
variability in soybeans. Agronomy Journal 47:314–318.
Khaliq, I., Noorka, L. R. and Khaliq, R. 2009. Estimation of
heritability and genetic advance for some quantitative characters in spring
wheat. Int. J. Agric. App. Sci. 1:76–78.
Kishore, S. N., Srinivas, T., Nagabhushanam, U., Pallavi, M.,
and Sameera, S. 2015. Genetic Variability, Correlation and Path Analysis for Yield and
Yield Components in Promising Rice (Oryza sativa L.)
Genotypes. SAARC J. Agri
13.1:99–108.
Maji, A. T., Shaibu, A. A., and
Cereals, N. 2012. Application of principal component
analysis for rice germplasm characterization and
evaluation. Journal of Plant Breeding and Crop Science 4.6:87–93.
https://doi.org/10.5897/JPBCS11.093
Moosavi, M., Ranjbar, G., Zarrini, H. N., and Gilani, A. 2015. Correlation between
morphological and physiological traits and path analysis of grain yield in rice
genotypes under Khuzestan conditions. Biological Forum – An
International Journal 7.1:43–47.
Mulugeta, S., Sentayehu, A. and Kassahun, B. 2012. Genetic Variability, Heritability, Correlation Coefficient and Path
Analysis for Yield and Yield Related Traits in Upland Rice (Oryza.
sativa L.). Journal of Plant Science 7.1:13–22.
Pachauri, A. K., Sarawgi, A. K., Bhandarkar, S., and Ojha, G. C.
2017. Agro-morphological characterization and morphological
based genetic diversity analysis of Rice (Oryza
sativa L.) germplasm. Journal of Pharmacognosy and Phytochemistry
6.6:75–80.
Panse, V. G. and Sukhatme, P. V. 1957. Genectics and Quantitative 1407 characters in relation to plant breeding.
Indian J. Genetics. 17:312-328.
Prasad, G.S., Sujatha, M., Subbarao, L. V. and Chaitanya, U.
2013. Studies on variability, heritability and
genetic advance for quantitative characters in rice (Oryza
sativa L.). Annals of Bio. Res. 4.6:372–375.
Rakesh, K. D., Sarawgi, A. K., Akash,
S. and Jitendra,
K. T. 2015. Genetic Variability Analysis for Various Yield Attributing and
Genetic Variability Analysis for Various Yield Attributing and Quality Traits
in Rice (O. sativa L.). The Bioscan an
International Quarterly Journal of Life Science 8.4:1403–1407.
Retrieved from www.bioscan.in
Robinson, H. F., Comstock, R. E. and Harvey,
P. H. 1949. (1949) Estimates of heritability and the
degree of dominance in corn. Agronomy Journal 41.8:353–359.
Saeed, S., Zia, M., and Ali, F. 2018. Genetic Variability, Correlation and Cluster Analysis in Elite
Lines of Rice. Journal of Scientific Agriculture 2:85–91.
https://doi.org/10.25081/jsa.2018.v2.900
Samadia, D. K. 2005. “Genetic variability studies in Lasora (Cordia myxa Roxb.).” Indian Journal
of Plant Genetic Resources 18.3:236–240.
Sameera, S., T. Srinivas, T., Rajesh, A.
P., Jayalakshmi, V., and Nirmala,
A. P. J. 2016. Variability and path co-efficient for yield and yield
components in rice s. Bangladesh J. Agril. Res.
41.2:259–271.
SAS (computer program). Version 9.4. Cary, NC: SAS Institute Inc;
2018.
Seyoum, M., Alamerew, S. and Bantte, K. 2012. Genetic Variability, Heritability, Correlation Coefficient and Path
Analysis for Yield and Yield Related Traits in Upland Rice (Oryza
sativa L.). Journal of Plant Sciences 7.1:13-22.
Singh, A., Singh, B., Panda, K., Rai,
V. P., Singh, A. K., Singh, S. P., Chouhan, S. K., Rai, V., Singh, P. K., & Singh, N. K. 2013. Wild rices of Eastern Indo-Gangetic
plains of India constitute two sub-populations harbouring
rich genetic diversity. Plant Omics 6.2:121–127.
Singh, T. V. J., Mohan, Y. C., and Raju,
C. H. D. 2019. Character association and path coefficient studies in rice
(Oryza sativa L.). Journal of Pharmacognosy
and Phytochemistry 8.3:383–387.
Subbaiah, P. V., Sekhar, M. R., Reddy, K. H. P. and Reddy, N. P. E. 2011.
Variability and genetic parameters for grain yield and its components and kerenel quality attributes in CMS based rice hybrids (Oryza sativa L.). International
Journal of Applied Biology and Pharamaceutical
Technology. 2.3:603–609.
Tuhina-khatun, M., Hanafi, M. M., Yusop, M. R., Wong, M. Y., Salleh,
F. M., and Ferdous, J. 2015. Genetic Variation, Heritability, and Diversity Analysis of Upland
Rice (Oryza sativa L.) Genotypes Based on
Quantitative Traits. Hindawi Publishing
Corporation, BioMed Research International 2015:1–8.
Retrieved from http://dx.doi.org/10.1155/2015/290861
Worede, F., Sreewongchai, T., Phumichai, C. and Sripichitt,
P. 2014. Multivariate analysis of genetic diversity among
some rice genotypes using morpho-agronomic traits.
Journal of Plant Sciences 9:14–24.


