INTRODUCTION
Beans are the most important grain
legumes for human consumption in the world. Beans represents one of the total
world production of pulse (19.3 Mt/ year; Norena-Ramirez et al., 2014).) and total production exceeds 23 million metric
tonnes (MT) of which 7 million MT are produced in Latin America and central
Africa where it is the staple food for many people due to its energy, protein,
dietary fiber and minerals content ( Norena-Ramirez et al., 2014). Beans come in many varieties of shapes, sizes and
colours, from pinto to pink, black and white, interesting enough, despite this
diversity in colour and size, the wild and domestic beans belong to the same
species, as do all of the colourful varieties of beans, which are believed to
be the result of a mixture of population bottlenecks and purposeful selection (Lerner,
2009).
Fungi are the most
frequent pathogenic agent and represent the major threat to this crop since
they attack the root parts and destroy the proper functioning of the plant in
taking up water and other nutrients (Nana et
al., 2015). The diseases can be caused by a single soil – borne pathogens,
resulting in disease complexes (Nana et
al., 2015). Rhizoctonia solani,
Fusarium oxysporum, F. spp. Phaseoli, F. solani F. spp. Phaseoli, Macrophomina phaseoli, Sclerotium
rolfsii, Meloidogyne spp., Aspergillus spp. (Domijan et al., 2015;Nana et al., 2015 ) are among the major pathogens known to impact bean
production in many countries in Latin America and Africa. Seeds are the
vehicles of transmission of several fungi and frequently introduce new pathogen
in exempt areas, so that the integration between seed health and germination
tests is recommended to control seed transmitted diseases (Francisco and
Usberti, 2008).
Fungi are known to
produce one or more toxic secondary metabolites in bean seeds. The presence of
these toxins in foods and food products is a serious health hazard to consumers
(Betina, 2012). Seed borne fungi pathogens are the principal producers of
mycotoxins associated with fungal growth on crops in the field and in storage
(Betina, 2012). It is widely acknowledged that Aspergillus and Penicillium species
are the most important mycotoxin-producing fungi in tropical countries, seen
mostly among adults in rural populations with a poor level of nutrition for
whom common beans is the staple food. (Tulpule and Bhat, 2012).
Although there have
been a number of investigations related to the bean, little is known about the
identities and prevalence of Aspergillus
spp. on bean seeds, therefore this study sought to evaluate the identities and
prevalence of Aspergillus spp. on
bean seeds.
MATERIALS AND METHODS
Study Area: These study was done in Ihiala Local Government Area (L.G.A.),
Anambra State, located at latitudes 5.85°N and longitudes 6.85°E on the
Southeast part of Nigeria. Ihiala is predominantly a low lying region on the
elevational plain of Manu river with all parts at 146 meters above sea level
Ihiala has rainforest vegetation with two seasonal climatic conditions. There
are rainy season and dry season which is characterized by harmattan between
December and February. Ihiala is characterized by the annual double maxima
rainfall with a slight drop in August break. The annual total rainfall is about
1600 mm with relative humidity of 80% at dawn. Ihiala has minimum daily
temperature of 18°C, annual minimum and maximum temperature ranges are about
22°C and 34°C respectively (Jim et al., 2009).
Sample Collection: A total of ninety
samples of bean seeds were collected randomly, from different shops and open
markets in Ihiala Local Government Area (L.G.A.), Anambra State. Sampling was
performed manually from different bags and basins, such that the bean seeds
were collected from different parts of the bags and basins. The samples were
aseptically pooled and mixed properly and formed one cup of the bean seeds in
sterile nylon bag, then the bean seeds were taken for analysis. The samples
were carefully labeled and then kept in a disinfected cooler, to maintain its
temperature and stability of the number of the isolates. The samples were
transported to the laboratory for analysis.
Isolation of the Fungi Isolates: This was done using
the method of Suleiman and Omafe (2013). Each sample was shared into two
groups. First group was aseptically soaked into distilled water for 30 minutes,
and the second group was disinfected by soaking for 1 minute in 1% Sodium
hypochloride and washed three times with distilled water, and then soaked in
the distilled water for 30 minutes. A 0.1 ml aliquot from the first group was
plated on Sabouraud Dextrose Agar (SDA) containing
chloramphenicol antibiotics (0.05%). Seeds from the second group was placed
at the rate of 25 seeds Per Petri dish
containing 20 ml of SDA supplemented with chloramphenicol antibiotics
(0.05%).These were incubated at room temperature (30±2ºC) for 5 days. The fungi
obtained were aseptically sub cultured on SDA containing chloramphenicol
antibiotics (0.05%) and incubated at room temperature (30±2°C)
for 5 days.
Identification of Fungal Isolates: The fungal isolates
were identified to the genus/species level based on macroscopic, microscopic
and molecular characteristics of the isolates obtained from pure cultures
(Watanabe, 2002).
Prevalence of the
Isolates:
The occurrences of each identified isolate were carried by determining the
number of times the isolate occurred in the samples, and also estimated the
percentage of the occurrence using the formula below:
Statistical Analysis: The occurrences of the isolates were
presented in percentages, and pairwise comparison of occurrence each isolate
was carried out using student “T” test.
RESULTS
A total ninety (90)
samples of bean seeds were screened for the presence of Aspergillus species, out of these, 32(35.56%) were positive to Aspergillus species surface
contamination whereas 13 (14.44%) were positive to Aspergillus species for internal contamination as show in Table 1.
The occurrence of Aspergillus species
was seen most on Oloka bean (70%)
seed samples for surface contamination and in Sokoto bean (50%) seed samples
for internal contamination. Iron white bean (10%) seed samples showed least
occurrence of Aspergillus species for
surface contaminant. Honey bean (0.00%), Iron white (0.00%) bean and Soya bean
(0.00%) samples did not record any occurrence of Aspergillus species for internal contaminants.
The isolates were
characterized macroscopically using their initial and their final appearances
on Sabouraud Dextrose Agar (SDA), colour on the reverse side of the plate and
colouring of the mycelium as shown in Table 2. Isolates Y1, Y2, M, Q and R have
almost similar morphology, dark brown to black mycelia and belong to the group
of Aspergillus known as Negri
section. Isolates Y1 and Y2 were almost identical except the slight different
in their mycelia appearances. Isolate R was also similar to isolates Y1 and Y2
but differ slightly in its initial appearances on SDA, colour on the reverse
side of the plate and colour of the mycelium. Similarly, isolates Q and M
showed slight variations in their initial appearances and colours of the
reverse side of the plates, Isolate X showed clear variation from isolates Y1,
Y2, M, Q and R: and possessed distinct features of the group of Aspergillus known as flavi section. The
characteristics features of the hyphae, conidia, conidiophores, vesicle,
sterigmata and metulae covering were basically used for the microscopic
features of the fungal isolate as shown in tables isolates Y1, Y2, M, Q and R
had almost similar microscopic features of Negri section group of Aspergillus. The five isolates differ in
the diameter of their vesicles: with slight variation in the colour of their
conidia. Isolate Q showed clear distinction of being uniseriate whereas other
isolates were biseriate. Isolates Y1 and Y2 have similar features but differ
from isolates M, Q and R by having brownish condidiophore. Isolate X was
clearly different from isolates Y1, Y2,
M, Q and R. Isolate X showed short conidiophores, yellowish green conidia,
three- quarter metulae covering and columnaric conidia head, which are features
of flavi section of Aspergillus.
The qualities of
nucleic acids (DNA) extracted from isolates x, Y1, Y2, M, Q and R were within
the stipulated range (1.80-1.90). Purity of nucleic acids (DNA) was determined
by calculating the ratio of the absorbance A260/A280 as
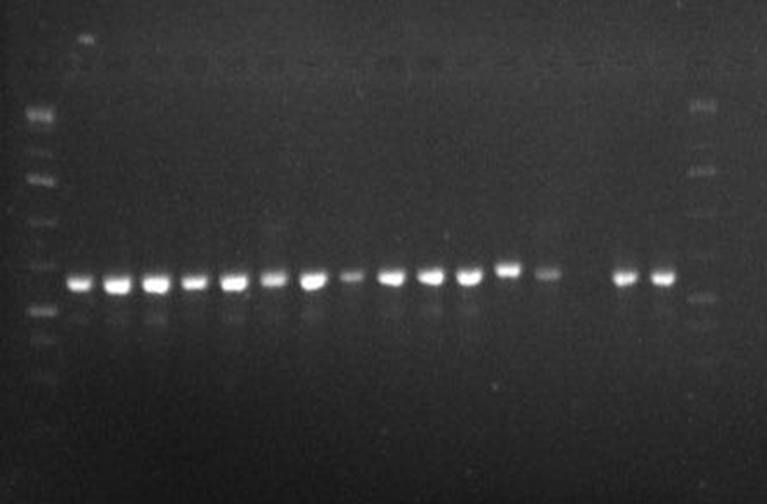
shown in Table 4. The gel characteristics of polymerase chain reaction (PCR) of
the nucleic acids (DNA) extracted from isolated X, Y1, Y2, M, Q and R is shown
in figure 1. The regions coding internal transcribed spacer rDNA (ITS1-1.58
ITS2), B-tubulin (Ben A) and Calmodulin (Cam) were amplified and
electrophoresed using 1.5% agarose. The photograph of the gel reviewed clear
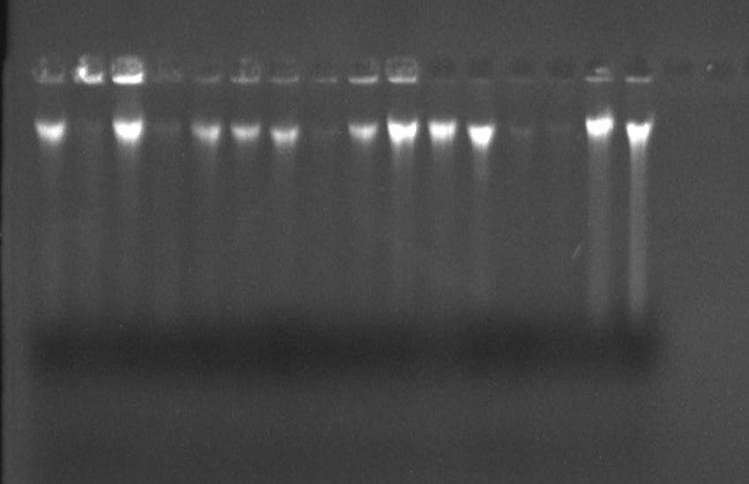
amplification of the selected regions for sequencing of the isolates. The
amplicons were cleaned (figure 2) and then sequenced. The amplicons generated
from nucleic acids (DNA) extracted from the isolates were cleaned and
re-electrophoresed as shown in figure 2. The sequencing of the amplified
regions of isolates X, Y1, Y2, M, Q and R showed 100% identities of each of the
isolates (Table 5).
The study revealed the
presence of Aspergillus flavus strain
Hus6 (isolates x), Aspergillus niger strain HG 48
(isolate Y1), Aspergillus niger
strain HUS 1 (Isolate Y2), Aspergillus
tubingiensis strain Em-CN1 (Isolate
M), Aspergillus aculeatus strains AN5
(Isolate Q) and Aspergillus awamori
strain DN-SN2 (Isolate R). Aspergillus niger strain Hus 1 was
mostly significant (P<0.05) in both surface and internal contamination of
the studies bean seed sample whereas Aspergillus
tubingiensis strain Em-CN1 and Aspergillus
aculeatus strain AN5 were not detected for internal contaminant. For
surface contaminants, Aspergillus
tubingiensis strain EM-CN1 (ATEM-CN1) A.
aculeatus strain AN5 (AA AN5) and A.
Awamori strains DA-SN2 (AWDA-SN2)
were not seen in Mesugar bean samples A.
flavus strain HUS 6 (AF HUS 6), ATEM-CN1 and AA AAN5 were not seen in Honey
bean samples, AF HUS6 and ATEM-CN1 were not seen in Potasco white bean sample, AA AN5 was not seen in Potasco brown bean samples, Aspergillus niger strain HG48 (ANHG48),
AFHUS 6 and ATEM-CN1 were not seen in Sokoto bean sample, ATEM-CN1 and AA An5
were not seen in Soya bean samples, and Aspergillus
niger strain HUS 1 (AN HUS1) was only seen in Iron white bean samples. For
internal contaminants, AN HUS1 was only seen in Mesugar, Potasco white
and Potasco brown bean samples,
ATEM-CN1 and AA AN5 were not seen in Oloka
and Sokoto bean samples, ANHUS1 and ANHG48 were only seen in Iron brown bean
sample, and no organism was seen in Honey, Iron white and Soya samples.
DISCUSSION
The presence of Aspergillus species in the studied been seed samples could be traced from poor
handling, management practices, transportation of the bean seed samples and
sanitary conditions distributed to the bean seed samples. Similar findings were
reported by many researchers (Immersed et
al., 2002; Maciorowski et al., 2007;
Iheukwumere et al., 2017). Many
studies have shown that bean seeds contaminated by pathogenic Aspergillus species could contribute to
human food-borne illness through the food-human chain. This shows that
preparation of bean seeds for human consumption requires microbiological safety
regulations to escape contamination by pathogenic microorganisms. Similar
deduction was drawn by different researchers (Davies and Wales, 2010; Chowdhuri
et al., 2011; Fredrick and Huda,
2011).
The variation in the
occurrence of Aspergillus species
from different types of been seeds observed in this study could be attribute to
the nature, texture, water activity and proportion of nutrients in the bean
seeds. Similar deductions were drawn by many researchers (Maciorowski et al., 2007; Iheukwumere et al., 2017).
The presence of Aspergillus flavus strain HUS6, Aspergillus niger strain HG48, Aspergillus niger strain HUS1 Aspergillus
tubingiensis strain EM-CN1, Aspergillus
aculeatus strain AN5 and Aspergillus
awamori strain DA-SN in the studied bean seed samples supported the
occurrence of Aspergillus species in
the bean seeds, and this corroborates with the report of many researchers
(Barakat, 2004; Damijan et al; 2014).
Traditionally, the laboratory defection of Aspergillus
species has relied on non-selective and selective enrichment media and
subsequent subculture on selective media. The introduction of molecular
techniques provides more sensitive and rapid technique for detecting these
fungi. The genes that code translation elongation factor-1; calmudulin,
β-tubulin and internal transcribed spacer rDNA region (ITS1-1.585-ITS2)
were amplified and used as barcode for the fungal identification. These genes
were also reported by other researchers (Samson et al., 2007; Varga et al., 2007).
The highest counts of
Aspergillus niger
strain Hus 1 recorded in the studied bean seed samples could be attributed to
human activities during processing, transportation and the storage of the bean
seeds. Similar deduction was made by Banford and Adebanjo (2011).
CONCLUSION
From this study, it was observed that Aspergillus flavus strain HUS6, Aspergillus niger strain H948, Aspergillus niger strain HUS1, Aspergillus tubingiensis strain EM-CNI, Aspergillus aculeatus strain AN5 and Aspergillus awamori strain DA-SN2 were
isolated from the studied bean samples, of which Aspergillus niger strain HUSI was the most predominant. This study
suggest proper handling and good hygienic measures must go in parallel with
good management practices to minimize the occurrence of the fungi in the
studied samples.
REFERENCES
Banford, S.A. and
Adebanjo, A. (2011). Mycotoxins in food in West
Africa. Current situation and possibilities of controlling it. African
Journal of Biotechnology 2:
254–263.
Bhat, R. V. and Miller, J. D. (2012).
Mycotoxins and food supply. International
Centre for Science and High Technology 2:
21–25.
Barakat,
R. (2004). Bacterial contamination of animal feed and its relationship to food
borne illness. Clinical Infection
Diseases 35: 859–865.
Betina,V.(2012).Mycotoxins and Environment. Prescott, Harley and
Klein’s Microbiology, Seventh Edition. McGraw Hill, New York, pp. 964–973.
Chowdhuri,
A., Iqbal, A., Giasuddin, M. and Bhuiyan, A. A. (2011). Study on isolation and
identification of Salmonella and Escherichia coli from different poultry
feed of Savar region of Dhaka, Bangladesh. Journal
of Science Resources 3(2):
403–411.
Damijam, J.
G. (2015). Pathological changes in, lungs and liver of mice in
immunocompromised individual following intramuscular administration
hydrocortisone acetate. Journal of
Science 38: 5–10.
Davies,
R. H. and Wales, A. D. (2010). Investigation into Salmonella contamination in poultry feed mills in the United
Kingdom. Journal of Applied Microbiology 109: 1430 –1440.
Domijan, A.M., Peracia, M., Zlender,
V. and Ivic. D. (2015). Seed borne fungi and ochratoxin A Contamination of Dry
Beans Food and Chemical Toxicdogy. Elsevier Limited, Amsterdam, Nether lands, pp.
427–432.
Francisco, F.G. and Usberti, R. (2008).
Seed health of common beans stored at constant moisture and temperature. Science Agriculture 65: 613 – 619.
Frederick,
A. and Huda, N. (2011). Salmonellas, poultry
house environments and feeds: A review. Journal
of Animal and Veterinary Advances 10(5):
679–685.
Iheukwumere,
I.H., Ejike, C.E. and Okeke, C.E. (2017). A trial to prevent sorbitol negative Escherichia coli infections in chicks using autogenous bacterin and
probiotics. Journal of Natural Science Research 7(4): 56–63.
Immerseel,
F. V., Cauwerts, K., Devriese, L. A., Haesebrouck, F. and Ducatelle, R. (2002).
Feed additives to control Salmonella in
poultry. World’s Poultry Science 58: 431– 443.
Jim, E., Cutler, A., George, S., Deepe, J. and Bruce,
S. (2009). Advance in combating fungal diseases. Klein Nature Reviews Microbiology 5: 13–28.
Lerner, D. (2009). Food and Indoor
Fungi. Second Edition, CBS Laboratory Manual Series Publishers, Utrecht, Netherlands, pp. 390–395.
Maciorowski,
K. G., Herrera, P., Kundinger, M. M. and Ricke, S. C. (2007). Animal feed
production and contamination by food borne Salmonella.
Journal of Consumer Protection and Food Safety 1:197–209.
Mohammed,
S. U., Anup, K., Anil, K. R. and Shipra, S. (2011). Identification of Escherichia coli through analysis of 16s
rRNA and 16s-23s rRNA internal transcribed spacer region sequences. Bioinformation 6: 370–371.
Nana, W.L., Eke, P., Atogho, T.B.,
Toghueo, K.R.M., Chatue, C.G., Pokaa, N.A., Ekounda, T.V., Etoa, F.X. and
Fekam, B.F. (2015). In-vitro fungitoxic
effects of cold and hot water extracts of three Ocimum species of species of fungi causing common bean (Phaseolus vulgaris L.) Root rot in
Cameroon. International Journal of
Current Microbiology and Applied Sciences 4: 596 – 606.
Norena-Ramirez, D.A.,
Valasquez-Ballesteros, O.J., Murillo-Perea, E., Mendez-Arteaga, J.J.,
Solanilla-Duque, J.F., and Murillo-Arango, W. (2014). Assessment of Phaseolus vulgaris L. and Vigna unguiculata L. Walp leaves for
antifungal metabolites against two bean fungal pathogens Colletotricum lindemuthianum and Phaeoisariopsis griseola. African
Journal of Biotechnology 13: 1657 – 1665.
Samson,
R.A., Hoekstra, E.S., Frisval, J.C. and Filtenborg, O. (2007). Introduction of
Food and Airborne Fungi, Seventh Edition. Delft, Netherlands pp. 91–99.
Suleiman
and Omafe, K. A. (2013). Identification and characterization of hyphomycetous
fungi in bean seeds. American Journal of pathology 114: 23–27.
Tulpule,
R.D. and Blat, C.C. (2012). Bean
anthracnose fact sheet. Department of Plant pathology, New York State
Agricultural Experiment Station, Geneva Cornell University, pp. 729–740.
Varga,J., Peteri, Z., Tabori, K., Teren, J. and Vagwolgyi, C.
(2007). Degradation of Ochratoxin A and other mycotoxins by Rhizopus isolates. International Journal of Food
Microbiology 99: 321–328.
Watanabe,
T. (2002). Pictorial atlas of soil and seed fungi. Morphologies of Cultured Fungi
and Key to Species. Lewis Publishers, London, Washington D.C., pp. 410–421.