By Yanda, FE; Tryphone,
GM (2023).
Microsatellite
Analysis of Common Bean (Phaseolus vulgaris
L.) Genotypes in Tanzania for Diversity in Seed Iron and Zinc
Micronutrients
Yanda, Focus Edson1;
Tryphone, George Muhamba2*
1Ministry of Agriculture, Manyoni District Council, P.O. Box 60, Manyoni,
Tanzania.
2Department of Crop Science
and Horticulture, College of Agriculture, Sokoine
University of Agriculture, P. O. Box 3005, Chuo Kikuu, Morogoro, Tanzania.
|
ARTICLE INFO
|
ABSTRACT
|
|
Article No.: 011723005
Type: Research
Full Text: PDF,
HTML,
PHP,
EPUB
DOI: 10.5281/zenodo.7655383
|
Common bean (Phaseolus vulgaris
L.) grain has big dietary role in supplying protein, energy, vitamins,
dietary fibre and micronutrients for millions of people worldwide. In
Tanzania, reports have shown existence of Fe and Zn micronutrient variabilities among the common bean genotypes.
Eighty-four (84) genotypes were collected to study such variability by seed
biochemical and leaf molecular marker analysis. The analyses revealed
significant (p < 0.001) genetic variability for seed Zn and Fe nutrient
contents. Highest seed Fe content was 118 ppm for genotype Kashiransoni and for seed Zn was 51.81 ppm for Imponzo 5 genotype (both were collected from
NPGRC). With seven (7) SSR primers associated with Fe and Zn traits, genetic
diversity was evaluated. A marker BM154 scored PIC value 0.967 mean while
marker BM160 had a lowest PIC score of 0.899. Using an estimated genetic
similarity value, two main clusters with sub clusters in the dendrogram were developed. To corroborate the UPGMA
analysis results, a Principal Coordinate Analysis (PCoA)
was performed and displayed outputs into scattered plot presentation. For Fe
micronutrient improvement purposes, Inula was
proposed to be crossed with Kashiransoni
while Roba with Imponzo
5 for Zn micronutrient improvement. Selection was based on both concentrations
of nutrients for each genotype and their genetic similarity distances.
|
|
Accepted: 20/01/2023
Published: 15/02/2023
|
|
*Corresponding Author
George Muhamba Tryphone
E-mail: muhatry@ gmail.com
|
|
Keywords: SSR, Common bean, Parent
selection, Iron and Zinc micronutrients, diversity
|
|
|
|
1.0 INTRODUCTION
Common bean (Phaseolus vulgaris
L.) is one of the principal foods and cash crop legumes grown in the tropics
and most of the production takes place in developing countries (Hillocks et al. 2006). Common bean grains have
big dietary role in supplying protein, energy, dietary fiber and micronutrients
for millions of people worldwide although its intake, in terms of nutrient
content level, does not satisfy their nutrient requirements (Hillocks et al. 2006). Although common beans are
grown largely by smallholder farmers for subsistence and mainly by women, about
40% of the total production from Africa is marketed (Wortmann
et al. 1998). It is estimated that
over 75% of rural households in Tanzania depend on common beans for daily
sustenance (CIAT, 2008).
Micronutrients like zinc and iron are among the important
nutrients required in small amounts in the diet to fulfill the nutritional
needs of people. Both of these micronutrients are critical to human well-being
and an adequate supply of iron and zinc helps to prevent iron deficiency which
may lead to reduced oxygen transportation in blood, immune function and
consequently to anemia (Blair et al. 2009).
Zinc deficiency may lead to reduced immune function, fetal brain cell
development and child’s growth and cognitive development as well as other
health problems of the developing world (Black, 2003). Zinc is also an
anti-cancer element because a potential anti-oxidant. Identifying common bean
with high level of these nutrients and integrating them in daily diet will help
to improve the situation.
In Tanzania, Tryphone and Nchimbi-Msolla
(2010) reported a large variability among common bean genotypes for Fe and Zn
nutrient concentrations. Such variability is influenced by genotype
environmental interaction and suggested improving climate adapted genotypes has
greater potential than improving common bean based on the difference in content
of Fe and Zn, for exploiting existing genetic diversity in Tanzania. Unfortunately,
there is limited information on how the common bean genotypes being adopted in
Tanzania vary genetically for iron and zinc. The knowledge of genetic diversity
patterns increases the efficiency for conservation, utilization and genetic
improvement of common beans (Beebe et al.
2000). Tremendous efforts have been made to delve deeper into the genetic
diversity of these traits in several crop species, including common bean
through the use of molecular markers (Talukder et al. 2010). Molecular markers used in
estimating the genetic diversity and level of heterozygosity
among plants and animals, thus have a great potential to help breeders develop
new and improved varieties (Kumar et al., 2008). Therefore, this study will
investigate the genetic diversity among germplasm
collection of common bean to obtain suitable parental genotype (s) based on
both genetic similarity distances and the nutrient levels of Fe and Zn.
2.0 MATERIALS AND METHOD
2.1 Location of the study
The study was conducted at Sokoine University of Agriculture (SUA) in Morogoro, Tanzania in the screen housed behind African seed
building located at latitude 6.84795 S and 37.65904 E at 543 m above the sea
level. The study was conducted for the period of December 2017 to October 2018.
2.2
Genotypes
Eighty-four (84) common
bean genotypes seed were collected from National Plant Genetic Resource Center
(NPGRC) at Arusha, Uyole
National Research Institute (UNRI) at Mbeya and SUA
at Morogoro. The seed was stored in a cold room
(12˚C) until planting at Sokoine University of
Agriculture, Department of Crop Science and Horticulture. Genotypes collected
were diverse, representing a range of seed types, ranging from different seed
coat colors, size and shape. During accessioning, there were no specific and
strict criteria in choosing a genotype to collect. The collected accessions are
as indicated in Table 1.
Table 1: Common bean (Phaseolus vulgarism L.) genotype accessions from
different regions of Tanzania
|
S/N
|
Accession number
|
Genotype
|
Classification
|
Seed Zn (ppm)
|
Seed Fe (ppm)
|
Origin
|
|
1
|
SUA222
|
Beti 10
|
Improved
|
34.37m-x
|
43.3d
|
Morogoro
|
|
2
|
NPGRC2154
|
Biliomunyungu
|
Landrace
|
34.65n-y
|
53.56l-p
|
Kagera
|
|
3
|
SUA501
|
Cal 143
|
Improved
|
31.1h-s
|
58.48r-u
|
Morogoro
|
|
4
|
UYL5011
|
CalimaUyole
|
Improved
|
36.69o-B
|
54.8m-q
|
Mbeya
|
|
5
|
SUA180
|
Canadian Wonder
|
Improved
|
34.17m-x
|
66.12A-E
|
Morogoro
|
|
6
|
SUA6301
|
Cheupe
|
Improved
|
43B-I
|
64.61z-B
|
Morogoro
|
|
7
|
NPGRC 286
|
Chilanda 6
|
Landrace
|
27.94d-m
|
53.04j-o
|
Rukwa
|
|
8
|
NPGRC 287
|
Chilanda 7
|
Landrace
|
34.1m-x
|
71.29G-I
|
Rukwa
|
|
9
|
NPGRC 134
|
Chilemba 4
|
Landrace
|
33.97m-x
|
44.29de
|
Rukwa
|
|
10
|
NPGRC 135
|
Chilemba 5
|
Landrace
|
31.86i-t
|
58.77s-v
|
Rukwa
|
|
11
|
NPGRC 306
|
Chilemba 6
|
Landrace
|
26.9c-j
|
61.68w-y
|
Rukwa
|
|
12
|
NPGRC 307
|
Chilemba 7
|
Landrace
|
25.75b-i
|
84.08PQ
|
Rukwa
|
|
13
|
NPGRC 133
|
Chilemba3
|
Landrace
|
42.91A-I
|
75.02K-M
|
Rukwa
|
|
14
|
NPGRC 401
|
Fibea
|
Improved
|
35.66o-z
|
57.55q-t
|
Morogoro
|
|
15
|
NPGRC 4312
|
FukamaOkole
|
Landrace
|
41.73z-I
|
53.26k-o
|
Kagera
|
|
16
|
NPGRC 3153
|
Gwezamenyo
|
Landrace
|
38.77u-E
|
61.7w-y
|
Kigoma
|
|
17
|
NPGRC 147
|
Ilanda / Kalinso
|
Landrace
|
38.12t-D
|
69.76F-H
|
Rukwa
|
|
18
|
NPGRC 334
|
Imponzo 4
|
Landrace
|
37.89t-D
|
64.13y-A
|
Mbeya
|
|
19
|
NPGRC 335
|
Imponzo 5
|
Landrace
|
47.89IJ
|
66.72A-E
|
Mbeya
|
|
20
|
NPGRC 337
|
Imponzo 7
|
Landrace
|
51.81J
|
72.4H-K
|
Mbeya
|
|
21
|
NPGRC 331
|
Imponzo1
|
Landrace
|
46.08G-H
|
55n-q
|
Mbeya
|
|
22
|
NPGRC 188
|
Imponzo8
|
Landrace
|
27.29d-l
|
90.57R
|
Mbeya
|
|
23
|
NPGRC 198
|
Imponzo9
|
Landrace
|
43.16C-I
|
76.57LM
|
Mbeya
|
|
24
|
NPGRC 4258
|
Inula
|
Landrace
|
20.5ab
|
32.36a
|
Kagera
|
|
25
|
SUA10
|
Jesca
|
Improved
|
25.97b-i
|
66.64A-E
|
Morogoro
|
|
26
|
NPGRC 70
|
Kablanketi
|
Landrace
|
30.84h-r
|
56.57q-t
|
Rukwa
|
|
27
|
NPGRC 3157
|
Kalambi
|
Landrace
|
42.71A-I
|
107.29T
|
Kigoma
|
|
28
|
NPGRC 2158
|
Kanyamunywa
|
Landrace
|
39.26v-F
|
52.46i-m
|
Kagera
|
|
29
|
NPGRC 3175
|
Kashiransoni
|
Landrace
|
45.93G-I
|
118.24U
|
Kigoma
|
|
30
|
NPGRC 69
|
Kasukanywele
|
Landrace
|
25.57b-i
|
65.05z-D
|
Rukwa
|
|
31
|
NPGRC 2190
|
Kibeho
|
Landrace
|
24.03a-f
|
68.83E-G
|
Kagera
|
|
32
|
SUA1400
|
Kigoma
|
Improved
|
32.45j-u
|
48.22fg
|
Morogoro
|
|
33
|
NPGRC 4336
|
Kiisiki
|
Landrace
|
24.72b-h
|
66.44A-E
|
Kagera
|
|
34
|
NPGRC 4265
|
Kisapuli
|
Landrace
|
28.89d-n
|
50.21g-j
|
Kagera
|
|
35
|
NPGRC 4259
|
Kya Karagwe
|
Landrace
|
34.77n-y
|
52.82j-o
|
Kagera
|
|
36
|
SUA444
|
Lyamungo 90
|
Improved
|
35.9n-z
|
65.94A-E
|
Morogoro
|
|
37
|
SUA333
|
Lyamungu 85
|
Improved
|
24.19a-g
|
55.7o-r
|
Morogoro
|
|
38
|
NPGRC 3511
|
Maharage - kienyeji
|
Landrace
|
23.9a-e
|
49.74g-i
|
Kigoma
|
|
39
|
NPGRC 3816
|
Maharage karanga
|
Landrace
|
33.53k-w
|
36.92b
|
Kigoma
|
|
40
|
SAU1300
|
Maini
|
Improved
|
34.36m-x
|
46.71ef
|
Morogoro
|
|
41
|
NPGRC 218
|
Malima
/ Ndondo
|
Landrace
|
44.78E-I
|
67.79C-F
|
Mbeya
|
|
42
|
NPGRC 4269
|
Maliwalinda
|
Landrace
|
32.43j-u
|
93.63S
|
Kagera
|
|
43
|
NPGRC 3164
|
Mamesa
|
Landrace
|
30.68h-q
|
74.1J-L
|
Kigoma
|
|
44
|
NPGRC 3141
|
Mbuvamutwe
|
Landrace
|
34.04m-x
|
76.37LM
|
Kigoma
|
|
45
|
SUA808
|
Mkanamna
|
Improved
|
37.11q-C
|
68.37EF
|
Morogoro
|
|
46
|
SUA601
|
Msafiri
|
Improved
|
22.67a-d
|
52.02h-m
|
Morogoro
|
|
47
|
SUA1003
|
Mshindi
|
Improved
|
41.86z-I
|
59.25t-w
|
Morogoro
|
|
48
|
SUA16
|
Msolin
|
Improved
|
40.04w-G
|
50.86g-l
|
Morogoro
|
|
49
|
NPGRC 3120
|
Mulembegwa
|
Landrace
|
34.64n-y
|
85.44Q
|
Kigoma
|
|
50
|
NPGRC 3150
|
Mutsinga
|
Landrace
|
40.26x-G
|
67.91D-F
|
Kigoma
|
|
51
|
NPGRC 2178
|
Mwanamwana
|
Landrace
|
35.97o-z
|
81.93OP
|
Kagera
|
|
52
|
NPGRC 3155
|
Mwanja
|
Landrace
|
45.44F-I
|
66.22A-E
|
Kigoma
|
|
53
|
NPGRC 3119
|
Mwolo
-yellow
|
Landrace
|
41.96z-I
|
62.93x-z
|
Kigoma
|
|
54
|
SUA909
|
Nanavala
|
Improved
|
39.94w-G
|
49.77g-i
|
Morogoro
|
|
55
|
SUA800
|
Nanka
|
Improved
|
31.03h-s
|
56.25p-s
|
Morogoro
|
|
56
|
NPGRC 2213
|
Ndimila enkobe
|
Landrace
|
37.52s-D
|
88.09R
|
Kagera
|
|
57
|
NPGRC 3005
|
Njano ndefu
|
Landrace
|
36.52o-A
|
66.82A-F
|
Kigoma
|
|
58
|
UYL5010
|
Njano Uyole
|
Improved
|
41y-H
|
50.13g-j
|
Mbeya
|
|
59
|
NPGRC 3154
|
Nyamanza
|
Landrace
|
34.88n-y
|
73.49I-k
|
Kigoma
|
|
60
|
UYL5015
|
Nyeupe Uyole
|
Improved
|
33.16j-v
|
67.98D-E
|
Mbeya
|
|
61
|
UYL5018
|
Pasi
|
Improved
|
25.36b-h
|
52.99j-o
|
Mbeya
|
|
62
|
SAU1007
|
Pesa
|
Improved
|
33.35k-v
|
56.75q-t
|
Morogoro
|
|
63
|
SUA200
|
Roba
|
Improved
|
18.23a
|
58.7s-v
|
Morogoro
|
|
64
|
SUA1009
|
Rojo
|
Improved
|
37.42s-D
|
77.76MN
|
Morogoro
|
|
65
|
NPGRC 4248
|
Ruhondela
|
Landrace
|
28.5d-n
|
50.33g-k
|
Kagera
|
|
66
|
NPGRC 2220
|
Rukurulana
|
Landrace
|
34.06m-x
|
80.1NO
|
Kagera
|
|
67
|
NPGRC 4352
|
Ruvunja
|
Landrace
|
34.83n-y
|
67.18B-F
|
Kagera
|
|
68
|
SAU500
|
Selian
05
|
Improved
|
37p-C
|
64.86z-C
|
Morogoro
|
|
69
|
SUA777
|
Selian
06
|
Improved
|
30.28f-o
|
61.54v-y
|
Morogoro
|
|
70
|
SUA11
|
Selian
94
|
Improved
|
36.67o-B
|
48.23fg
|
Morogoro
|
|
71
|
NPGRC 3156
|
Seredi
|
Landrace
|
45.4F-I
|
51.89h-m
|
Kigoma
|
|
72
|
NPGRC 4221
|
Shona
|
Landrace
|
43.58D-I
|
76.88LM
|
Kagera
|
|
73
|
NPGRC 4322
|
Shona
egunia
|
Landrace
|
29.15e-n
|
48.01fg
|
Kagera
|
|
74
|
NPGRC 111
|
Soya
nano
|
Landrace
|
23.96a-f
|
60.7u-x
|
Morogoro
|
|
75
|
SUA1010
|
Sua
90
|
Improved
|
33.64k-w
|
49.15f-h
|
Morogoro
|
|
76
|
NPGRC 1604
|
Tichakuronza
|
Landrace
|
47.23H-J
|
72.14H-J
|
Kagera
|
|
77
|
NPGRC 3182
|
Ugweza
|
Landrace
|
33.68l-w
|
40.15c
|
Kigoma
|
|
78
|
UYL5017
|
Uyole
03
|
Improved
|
27.26d-k
|
57.3q-t
|
Mbeya
|
|
79
|
UYL5016
|
Uyole
04
|
Improved
|
37.27r-D
|
67.85C-F
|
Mbeya
|
|
80
|
UYL5012
|
Uyole
16
|
Improved
|
37.15q-C
|
67.26B-F
|
Mbeya
|
|
81
|
UYL5020
|
Uyole
94
|
Improved
|
33.21j-v
|
60.81u-x
|
Mbeya
|
|
82
|
UYL5013
|
Uyole 96
|
Improved
|
25.63b-i
|
38.35bc
|
Mbeya
|
|
83
|
UYL5009
|
Uyole84
|
Improved
|
20.68a-c
|
34.37a
|
Mbeya
|
|
84
|
SUA1001
|
Zawadi
|
Improved
|
30.47g-p
|
53.37l-o
|
Morogoro
|
|
|
|
Mean
|
|
34.44
|
62.533
|
|
|
|
|
SEM
|
|
1.845
|
0.898
|
|
|
|
|
%CV
|
|
9.3
|
2.5
|
|
|
|
|
P-value
|
|
<.001
|
<.001
|
|
Key:
UYL – Uyole, SUA - Sokoine University of Agriculture, NPGRC- National Plant
Genetic Resource Center, SEM- Standard error of means and %CV- Percentage
coefficient of variation
2.3 Soil sampling and chemical
analysis
Composite soil samples obtained were analysed
as described by Carter (1993). Bulk soil samples were taken at a depth of 0 -
20 cm on an area of 2 × 2 m2. Composite soil constituted nine
sub-samples randomly collected from forestry area covering 1.0 ha. For pot
experimentation, sub-samples were thoroughly mixed, sterilized, air dried and
ground to pass through an 8.0 mm mesh. Further preparations for soil analysis
were carried whereby, the 2.0 mm sieved composite soil samples were used for
laboratory physical and chemical analyses. Composite soil samples obtained were
analysed for pH, cation
exchange capacity, exchangeable bases (Ca, K, Mg and
Na), micronutrients (Fe, Zn, Mn and Cu), Nitrogen
(N), available phosphorus, particle size distribution and organic carbon (OC)
as described by Carter (1993). The soil pH was determined in water at a soil: water
ratio of 1:2.5 suspension using pH meter. Electrical conductivity was measured
in 1:2.5 soil: water using the electric conductivity
meter (Thomas, 1996). Cation exchange capacity (CEC)
was determined by the ammonium-acetate saturation method and quantification of
exchangeable bases: K, Ca, Na and Mg were determined
from the ammonium-acetate filtrates following the Lindsay and Norvell (1978) methods. Exchangeable calcium (Ca) and magnesium (Mg) were determined by atomic absorption
spectrophotometry whereas K and Na were extracted using ammonium acetate and analysed by flame spectrophotometry. The DTPA extractable
Cu, Fe, Mn and Zn were determined by atomic
absorption spectrophotometry (Lindsay and Norvell,
1978). Total nitrogen was determined by the micro-Kjeldahl
digestion distillation method (Bremner and Mulvaney, 1982). Soil extractable P was determined by using
the Bray-1-P method by Kuo, (1996) and colour was developed by the ascorbic acid-molybdate blue method (Murphy and Rilley,
1962). Particle size analysis was determined by the hydrometer method after
dispersing the soil samples with sodium hexametaphosphate
solution (Gee and Bauder, 1986). Soil textural
classes were determined using the USDA textural class triangle (USDA, 1975).
2.4
Screen house experimentation and agronomical practices
Eighty-four (84) genotypes
were arranged in a completely randomized design (CRD) and replicated three times.
Before sowing, the 4 kg potted soil was watered and allowed to stay for one
day. Four seeds were sown; thinning was done at age of 10 days after emergence.
Irrigation by re-introducing trapped water (infiltrates) on bottomed trays was
carried out regularly to maintain the moisture content.
2.5
Chemical analysis for iron and zinc nutrients in common bean seeds
After harvesting, bean
seeds from each pod of individual plant were mixed thoroughly and taken for
chemical analysis. Dry ash method of extraction was undertaken to determine
iron and zinc content (Jorhem, 1993). Digested plant
samples were taken for reading absorbance values using the Atomic Absorption
Spectrophotometer (AAS) method. A standard curve was computed and used to
determine the concentration of each sample using Microsoft excel program
(version 2016).
2.6
Genomic DNA extraction
Total genomic DNA was
extracted from a trifoliate leaf tissue sample of 14 days old plants by using
the Quick-DNA Plant/Seed Mini-Prep Kit protocols. DNA quality and quantity were
assessed on gel electrophoresis stained with safe view DNA loading dye
(EZ-vision) EZ vision. Seven simple sequence repeat (SSR) markers were used to
detect the polymorphisms and assess genetic diversity of 84 common bean
genotypes collected from two organizations, with several sources of their
origin and marker diversity as shown in Table 1and 3 respectively.
2.7
SSR markers and PCR amplification
SSR primer pairs were
selected on the basis of the published genetic diversity analysis on common
bean by (Talukder et
al. 2010). Seven microsatellite primers with high polymorphism levels
(≥0.5) were selected and used for final polymerase chain reaction (PCR)
amplification (Table 10) for investigating the genetic diversity on Fe and Zn
micronutrients in the dry bean accession. The DNA amplification was carried out
according Talukder et al. (2010) protocol with minor
modifications. A final volume of 25 μl for the
PCR reaction mix included 1.5 μl genomic DNA,
0.1 μl of forward and reverse primers, 12.5 μl of 2X× TaqMaster mix and
10 μl of nucleotide free sterile double
distilled water. The temperature profile in the thermocycler
was of one cycle of 95 0C for 3 min; 40 cycles of 95 0C for 1 min,
from 55 to 57 0C for 1 min, and 72 0C for 2 min; and one
cycle of 72 0C for 10 min.
2.8
Electrophoretic separation and visualization of amplicons
Superfine agarose (2 %) gel preparation involved the addition of 150
ml of chilled 1x TBE buffer (pH 8.0) to a flask with 3g of superfine agarose powder. Twenty microlitres
(20μl) of safe view DNA loading dye (EZ-vision) was added to the gel and
cast on a tray to set at room temperature. A gel was then carefully placed in a
tank with plenty of 1x TBE buffer and into each well, 3 μl
of the samples were carefully loaded with the first well having 1 μl of DNA ladder (100 bp).
Gel electrophoresis was run for 200 min at 120 volts. The bands were visualized
under an ultra violet trans-illuminator and photo-shopped using the Picasa
software.
2.9
Data analysis
2.9.1 Chemical analysis
Obtained
absorbance values were processed using MS excel (2018)
to determine the concentrations through R square value by establishment of
standard curve. The treatment effects were analyzed by analysis of variance (ANOVA)
using GenStat V. 25 software. In additional, Fisher’s
least significant difference (LSD) was used for mean separation at 5% level of
probability.
2.9.2 Genetic diversity
Gel images obtained after
electrophoresis were processed using IMAGE J software to determine the
molecular weight of the SSR marker bands in reference to the loaded DNA ladder.
The determined molecular weights of the SSR marker bands were analyzed by
POWERMARKER V.3.25 for estimating the mean number of observable alleles (Na),
major allele frequency, allele frequency, observed heterozygosity,
expected heterozygosity, gene diversity and the
polymorphic information content (PIC).
2.9.3 Cluster analysis
Genetic diversity analyses
were conducted using numerical taxonomy and the multivariate analysis system, NTSYSpc V. 2.2. Genetic similarity values were computed
between all possible pairs with the SIMQUAL option and ordered in a similarity
matrix. The similarity matrix was run by sequential, agglomerative,
hierarchical, nested (SAHN) clustering with the unweighted
pair group with arithmetic averaging (UPGMA) method as an option. The dendrogram and cluster groupings were constructed by the
UPGMA clustering algorithm from the SAHN option of NTSYSpc
v.2.2.
2.9.4 Principal coordinate analysis
To corroborate the
interrelationships between the common bean genotypes, a scatter plot
representation was built using principal coordinate analysis (PCoA) with the Nei’s similarity
coefficient. The analyses were performed using GenAlEx
statistical software V 6.5 (Peakall and Smouse, 2012).
3.0 RESULTS
3.1
Soil chemical analysis
The experimental forestry
soils had medium to high chemical and sandy clay loam textural class as
physical characteristics (Table 2). The analysed
composed forestry soil based on the selected soil parameters, showed optimal
condition that favour growth of common bean as
described by Landon, (1991). Therefore, the forestry soils were suitable for
production of common beans and other field crops like cereals.
Table
2: Physical-chemical properties of the experimental forestry soils
|
Soil
parameter
|
Values
|
Remark
(Landon,
1991).
|
|
pH
in water
|
6.79
|
Neutral
|
|
Electrical
Conductivity (EC) (µS /cm)
|
451
|
Medium
|
|
Cationic
Exchange Capacity (CEC)
|
32.6
|
High
|
|
Organic
Carbon (% OC)
|
2.62
|
High
|
|
Organic
matter (% OM)
|
4.52
|
High
|
|
Nitrogen
(%)
|
0.50
|
Medium
|
|
Carbon:
Nitrogen ratio
|
9.04
|
Good
quality of the OM
|
|
Phosphorous
(mgkg-1)
|
9.33
|
Medium
|
|
Extractable
K (Cmol (+) kg-1)
|
1.90
|
High
|
|
Extractable
Na (Cmol (+) kg-1)
|
0.14
|
Low
|
|
Extractable
Mg (Cmol (+) kg-1)
|
0.41
|
Low
|
|
Extractable
Ca (Cmol (+) kg-1)
|
16.85
|
High
|
|
DTPA
Extractable micronutrients (mg kg-1)
|
|
|
|
Fe
|
34.96
|
High
|
|
Zn
|
4.08
|
High
|
|
Particle
size analysis (PSA)
|
|
|
|
%Clay
|
33.56
|
|
|
%Silt
|
9.64
|
|
|
%Sand
|
56.8
|
|
|
Textural
class
|
Sandy
clay loam (USDA, 1975)
|
3.2
Chemical analysis for iron and zinc contents on common bean seeds
The iron content in seed
for collected genotypes of Common bean differed significantly (P<0.001)
(Table 1). It varied from 32.36 to 118.24 ppm, with a mean of 62.533ppm.
Genotypes of Kashiransoni, Kalambi and Maliwalinda were observed with the highest seed
iron (Fe) contents of 118.24 ppm, 107.29 ppm and 93.63 ppm respectively. On the other hand, 32.36 ppm, 34.37 ppm and 36.92
ppm were the lowest contents of iron observed in genotypes of Inula, Uyole84 and Maharage-karanga respectively.
The genotypes differed significantly (P<0.001) in seed zinc contents (Table
1). Values varied from 18.23 to 51.81 ppm with a mean of 34.44 ppm. Genotypes Imponzo5, Imponzo7 and Tichakuronza were observed with the highest seed
zinc (Zn) contents of 51.81 ppm, 47.89 ppm
and 47.23 ppm respectively meanwhile 18.23 ppm, 20.5 ppm and 20.68 ppm were the
lowest contents of zinc observed in genotypes Roba, Inula and Uyole 84 respectively.
3.3
Allelic diversity of common bean accessions
The key parameters used to
define genetic diversity among the common bean accessions from Tanzania are
presented in Table 3. There was a high polymorphism with a mean of 24.857
alleles per locus and a range of 17 to 40 alleles in the germplasm.
The frequency for the major allele ranged from 0.064 for primer BM154 to 0.208
for the primer BMd16 with mean of 0.139. In total, the 7 markers detected 174
alleles and PIC ranging from 0.892 for the primer BMd16 to 0.967 for the primer
BM154 with the mean of 0.923. The overall mean for the expected heterozygosity ranged from 0.899 for BMd16 primer to 0.967
for BM154 primer. The observed heterozygosity was
observed on the two primers of BM160 and BM181 with value 0.025 and 0.027
respectively.
Table
3: Genetic information for the markers used to detect the genetic diversity of
84 Common bean genotypes collected from different regions of Tanzania for the
Fe and Zn traits.
|
S/N
|
Marker
|
Sample
amplified
|
Major
allele frequency
|
Allele
number
|
Gene
Diversity
|
He
|
Ho
|
PIC
|
|
1
|
BM154
|
78
|
0.064
|
40
|
0.929
|
0.967
|
0.000
|
0.967
|
|
2
|
BM160
|
79
|
0.120
|
24
|
0.940
|
0.939
|
0.025
|
0.936
|
|
3
|
BM170
|
75
|
0.107
|
24
|
0.937
|
0.937
|
0.000
|
0.934
|
|
4
|
BM181
|
73
|
0.192
|
23
|
0.901
|
0.900
|
0.027
|
0.893
|
|
5
|
BM211
|
68
|
0.147
|
23
|
0.922
|
0.921
|
0.000
|
0.917
|
|
6
|
BMd16
|
72
|
0.208
|
17
|
0.899
|
0.899
|
0.000
|
0.892
|
|
7
|
BMd33
|
66
|
0.136
|
23
|
0.927
|
0.927
|
0.000
|
0.922
|
|
Mean
|
|
|
0.139
|
24.86
|
0.922
|
0.927
|
0.007
|
0.923
|
3.4
Cluster analysis
For
a better understanding of the genetic relationship among common bean genotypes,
the genetic similarity (GS) values were submitted to hierarchical clustering by
UPGMA. The dendrogram based on Nei’s
coefficient of genetic distance suggested the existence of two clusters with
sub clusters (Figure 1). The Cluster I was further subdivided into two sub
clusters of A and B in total comprised of 31 and 2 genotypes of common bean
respectively. Cluster II was also divided into two sub cluster designated as A
and B with a total of 28 and 23 genotypes respectively (Table 4).
Table
4: Distribution of genotypes to different clusters based on the UPGMA methods
|
Cluster
|
Sub
cluster
|
Number of genotypes
|
Name of genotypes
|
|
I
|
A
|
31
|
Jesca,
Kibego, Uyole 90, Sua 90, Kanyamunywa, Mshindi, NyeupeUyole, Uyole 94, Maharage-kienyeji, Zawadi, Shona-Egunia, Chilemba 4, Uyole 03, Rojo, Rukurulana, Kasukanywele, Chilemba 7, Biliomunyungu, Mwolo-Yellow, Maini, Msolin, Imponzo 9, Imponzo 1, Malima/Ndondo, Calima-Uyole, Fibea, Pasi, Msafiri, Kigoma, NjanoUyole and Njanondefu
|
|
B
|
2
|
Kablanketi
and Uyole 04
|
|
II
|
A
|
28
|
Selian
94, Ruvunja, FukamaOkole,
Ilanda/kalinso, Canadian
wonder, Uyole 16, Maharagekaranga,
Lyamungu 85, Cal 143, Inula,
Ruhondela, Chilemba 6, Imponzo 8, Kiisiki, Ugweza, Imponzo 7, NdimilaEnkobe, Nyamanzi, Chilemba 3, Mwanamwana, Maliwalinda, Mamesa, Mbuvamutwe, Mwanja, Mulembegwa, Seredi and Kashiransoni
|
|
B
|
23
|
Soya
Njano, Beti 10, Nanka, Roba, Lyamungu 90, Imponzo 5, Chilanda 7, Selian 06, Tichakuronza, Uyole 84, Mkanamna, Mutsinga, Shona, Selian 05, Kalambi, Chilemba 5, Chilanda 6, Kyakaragwe, Cheupe, Gwezamenyo, Nanayala, Pesa and Kisapuli.
|
3.5 Principal co-ordinate analysis
The common bean genotypes
were represented by a scatter plot in 2-dimensions using the results of the
principal coordinate analysis (PCoA) obtained by
estimating the Nei’s genetic similarity distance. It
revealed the global structure similar to the dendrogram
analysis, but the distribution of these accessions was shown more clearly in
scatter plot (Figure 2). Overall, the clustering pattern of the genotypes in
the principal coordinates analysis corresponds with the dendrogram
derived from UPGMA (Figure 1).






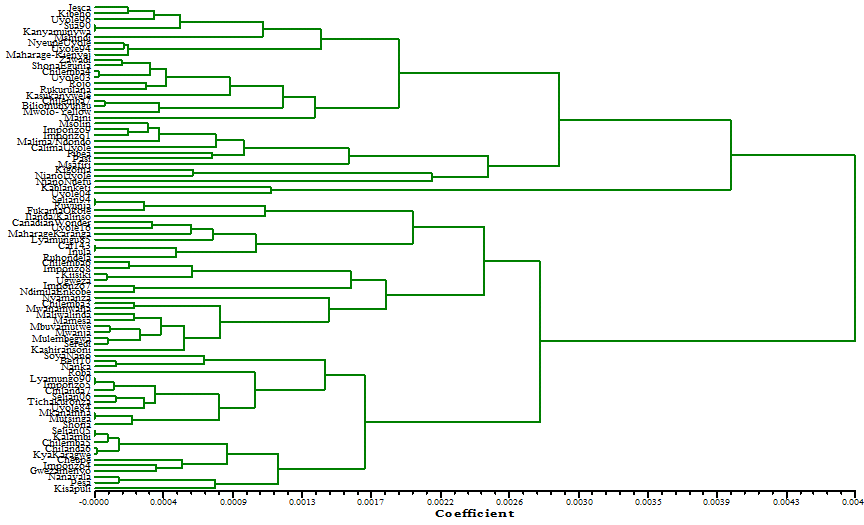
Figure 1: Dendrogram representing the genetic diversity among the 84
accessions of Common bean of Tanzania based on Dice coefficient
of
similarity matrix data using UPGMA cluster analysis

Figure 2: Principal
coordinate analysis (PCoA) of common bean accessions
from the microsatellite diversity based on molecular weight
(band sizes) of alleles.
4.0 DISCUSSION
4.1
Iron and zinc contents in common bean
The high seed Fe and Zn
content were expressed with genotypes of Kashiransoni and Imponzo5. Both genotypes were sourced at
the gene bank of Tanzania (NPGRC) being collected from Kigoma
(Kashiransoni)
and Mbeya (Imponzo
5) regions. There was no environmental contribution to the significantly
(p<0.001) variability of both seed Fe and Zn content on the experimented
genotypes of common bean because the study was conducted in screen-house
conditions hence differences in their genetic makeup could have resulted into
such variability.
4.2
Genetic diversity of common bean genotypes
The employed UPGMA
clustering method divided the common bean genotypes into two main clusters. The
main two branches of the dendrogram have showed a
considerable range of genetic diversity of genotypes in respect to the genetic
similarity distance being less than 9e-04, suggesting existence of low genetic
diversity based on Fe and Zn traits. This existence could be due to either the genotypes in equation comes from the similar origin or the
SSR markers used could not clearly detect large range of diversity. Similar
results for the diversity of common bean genotypes for iron and zinc nutrients
of Tanzania corresponds with numerous and deliberate efforts done by other
researchers worldwide by using various molecular techniques including SSR
marker. Talukder et
al. (2010) observed low level of genetic diversity because since the GS for
the common bean genotypes was more than 50%, hence
suggesting low genetic variability. Low genetic variability between the two
main clusters does hinder selection of parental genotypes for cross breeding
programs due to significant (p<0.001) difference in variabilities
of seed Fe and Zn content. Thus, it can be expected that the introgression of a
desired trait would be easier for this cross without sacrificing valuable
trait(s) and/or adding unwanted trait(s) as a consequence of linkage drag (Talukder et al. 2010).
4.3
Selection of genotypes for breeding programs
Selection of genotypes for
breeding purpose based on micronutrient variability and genetic diversity has
been suggested in many crop species including common bean by Talukder et al.
(2010) and rice (Gregorio, 2000). From the current results of genetic diversity
analysis, several breeding strategies can be derived; however, the interest at
present is to develop populations to map Zn and Fe content traits and to breed
common bean cultivars with enhanced Zn and Fe contents.
Based on seed Iron (Fe) and
Zinc (Zn) contents of the genotypes and genotypic variability at molecular
level, the cross-breeding program for seed Fe content was designed to cross the
genotypes with high Fe content namely Kashiransoni,
Kalambi and Maliwalinda
while low Fe content genotypes were Inula, Uyole84
and Maharage-karanga. The high Fe contents
genotypes Kashiransoni, Kalambi
and Maliwalinda belonged to clusters II-A,
II-B and II-A respectively and those of low Fe contents viz. Inula, Uyole 84 and Maharage-karanga belonged to clusters; II-A, II-B
and II-A respectively. The genetic similarity distance values of genotypes Inula with Kashiransoni,
Kalambi and Maliwalinda
are 1.704e-4, 0 and 1.0116e-3 respectively and those of Uyole 84 with Kashiransoni,
Kalambi and Maliwalinda
are 3.91e-3, 3.31e-3 and 3.20e-3 respectively
and those of Maharage-karanga with Kashiransoni, Kalambi
and Maliwalinda are 8.89e-4, 2.79e-3
and 1.03e-3 respectively. These values indicate that the selected
genotypes are divergent at the molecular level with significant differences in
seed Fe content and could result in better segregation and recombination of the
desired alleles in successive generations during population development.
For the crossing program to
enhance seed Zn content, genotype Imponzo 5
with high Zn content (ppm) was selected against the genotypes of Roba and Inula with low
zinc contents. The high Zn content genotype of Imponzo
5 belonged to clusters II-A and the low Zn content genotypes of Roba and Inula
belonged to clusters II-B and II-A respectively. The genetic similarity
distances value of genotype Roba with Imponzo 5 was 8.72e-4 and that of Inula with Imponzo 5
was 5.28e-4. These values indicate that the selected genotypes are
divergent at the molecular level with significant differences in seed Zn
content and could result in better segregation and recombination of the desired
alleles in successive generations during population development. Crossing
combinations with genotypes between and within cluster having high genetic
diversity and mineral content would be expected to accumulate positive alleles
derived from unique sources and generate breeding lines with even higher seed
mineral contents.
Also, crossing of the
selected genotypes would be of significant important for genetic studies
because they’re contrasting for the observed traits. Genetic studies offer an
opportunity of utilizing the genetic information like inheritance patterns of
Fe and Zn, additive effect and dominance effects of gene can be determined as knowledge
of heritability is crucial for decisions making concerning screening and
breeding methodologies and the scale of breeding.
5.0 CONCLUSION
The variability in iron and
zinc content was significant (p<0.001) across all experimented genotypes. The
range of seed iron content was 32.36 for Inula to
118.24 ppm for Kashiransoni genotype with mean
of 62.533 ppm. For seed zinc content it ranged from 18.23 ppm for Roba to 51.81 ppm for Imponzo
5 with a mean of 34.44 ppm. The SSR primers used in this study showed to be
effective on detecting the alleles bonded to the trait of iron and zinc. The
high PIC (0.967) value was shown by BM154 primer which detected 40 alleles for
78 samples of genotypes that were amplified. The overall results obtained by
SSR analysis of the common bean genotypes in the present study has shown that
there is genetic divergence among the collected genotypes based on the traits.
The UPGMA analysis together with the scattered plot presentation (PCoA) has successfully grouped the genotypes into sense of
similarity by using the estimated genetic similarity distance. It is recommended
that the genotypes Kashiransoni and Inula for iron and Roba
and Imponzo 5 for zinc as breeding
materials as sources Fe and Zn. Introgression
of gene for Fe and Zn enrichment should rely on consumer preferred common bean
genotypes as a key group during participatory plant breeding (PPB) programs. Further,
the inheritance patterns of Fe and Zn traits has to be studied since the
knowledge of heritability is crucial for decisions making concerning screening
and breeding methodologies and the scale of breeding.
REFERENCES
Beebe S, Sckroch
PW, Tohme J, Duque M C, Pedraza
F. Nienhuis J.
2000. Structure of genetic diversity among common bean
landraces of Middle American origin based on correspondence analysis of RAPD.
Crop Science, 40: 264-273.
Black R E. 2003. Zinc deficiency,
infectious disease and mortality in the developing world. Journal of Nutrition, 133 (5): 1485-1489.
Blair MW, Astudillo
C, Grusak M A, Graham R, Beebe S E. 2009. Inheritance
of seed iron and zinc concentrations in common bean (Phaseolus
vulgaris L.). Molecular breeding,
23:197-207.
Bremner J M. Mulvaney C S. 1982.
Total nitrogen. In: Methods of Soil Analysis. Part 2, Agronomy Monograph number. 9. (Edited by Page L A,
Miller R H. Keeney D R.). American
Society of Agronomy (pp.
595-624). Madison Wisconsin, USA.
Carter M. 1993. Soil sampling and methods of analysis.
(Editor). Lewis Publishers, CRC Press, Inc., Boca
Raton, FL. ISBN: 0-87371-861-5.
CIAT (International Center for Tropical
Agriculture). 2008. The impact of improved bean production technologies in
Northern Tanzania. Highlights of CIAT in Africa. Retrieved July 23rd 2018 from
http://www.ciat.cgiar.org/work/Africa/Documents/highlight42.pdf
Gee G W.
Bauder J W. 1986. Particle-size analysis.
In: Methods of Soil Analysis, Part 1. Physical and
Mineralogical Methods. Agronomy series number 9. (Edited
by Klute A). American Society of Agronomy (pp. 383-409). Madison Wisconsin, USA.
Gregorio G B. Senadhira
D. Htut T. Graham R D. 2000. Breeding
for trace mineral density in rice. Food
and Nutrition Bulletin, 21: 382-386.
Hillocks R J. Madata C S. Chirwa R. Minja E M. Nchimbi-Msolla S. 2006. Phaseolus bean improvement in
Tanzania, 1959–2005,” Euphytica, 150. (1-2): 215–231. DOI:10.1155/2013/436064.
Jorhem L. 1993. Determination of
metals in foodstuffs by atomic absorption spectrophotometry after dry ashing: NMKL inter-laboratory study of lead, cadmium, zinc,
iron, chromium and nickel. Journal of the
Association of Analytical Communities International, 76:708-813.
Kumar V. Sharma S. Kero
S. Sharma S. Sharma K. Kumar M. Bhat K V. 2008. Assessment of genetic diversity in common
bean (Phaseolus vulgaris L.) germplasm
using amplified fragment length polymorphism (AFLP). Science of Horticulture, 116:138-143.
Kuo S. 1996. Phosphorus. In: Methods of Soil Analysis, Part 3. Chemical Methods. Soil
Science Society of America (pp. 869 – 920). Madison Wisconsin, USA.
Landon J R. 1991. Booker Tropical Soil Manual. A handbook of soil survey and agricultural land evaluation in the
tropics and sub-tropics. Longman Scientific and
Technical /Booker Tate Ltd., London. p. 474.
Lindsay W L.
Norvell W A. 1978. Development
of DTPA soil test for zinc, iron, manganese and copper. Soil Science Society of American Journal,
42:421-428.
Murphy I. Rilley J P. 1962. A modified single solution method for determination of phosphate in
natural water. Anal. Chem. Acta
27: 31-36. Nutrition, 26(11):
2307-2319.
Nchimbi-Msolla S. Tryphone G M. 2010.
The Effects of the Environment on Iron and Zinc
Concentrations and Performance of Common Bean (Phaseolus vulgaris L.) Genotypes. Asian Journal of Plant Sciences, 9:
455-462. DOI: 10.3923/ajps.2010.455.462.
Peakall R. Smouse P E. 2012. GenAlEx 6.5: Genetic analysis in Excel. Population
genetic software for teaching and research-an update. Bioinformatics. 28(19): 2537-9.
Talukder Z
I. Anderson E. Miklas P N. Blair M W. Osorno J. Dilawari M. Hossain K G. 2010.
Genetic diversity and selection of genotypes to enhance Zn and Fe content in
common bean. Canadian Journal of Plant
Science, 90: 49-60.
Thomas G W. 1996. Soil pH
and Soil Acidity. In: Methods of Analysis, Part 3. Chemical.
Methods. American
Society of Agronomy (pp. 475 – 490). Madison Wisconsin, USA.
Tryphone G M. Nchimbi-Msolla S.
2010. Diversity of common bean (Phaseolus
vulgaris L.) genotypes in iron and zinc contents under screen house
conditions. African Journal of
Agricultural Research, 5(8):746.
USDA. (1975).
A Basic System of Soil Classification for Making and Interpreting Soil Surveys.
Soil survey staff, Soil Conservation Service (pp.754),
Washington DC.
Wortmann C S. Kirkby
R A. Eledu C A.
Allen D J. 1998. Atlas of Common Bean inAfrica. CIAT, Cali, Colombia.
|
Cite this Article: Yanda, FE; Tryphone, GM (2023). Microsatellite Analysis of Common
Bean (Phaseolus vulgaris L.) Genotypes in Tanzania
for Diversity in Seed Iron and Zinc Micronutrients. Greener
Journal of Plant breeding and Crop Science, 11(1): 1-12. https://doi.org/10.5281/zenodo.7655383
|


